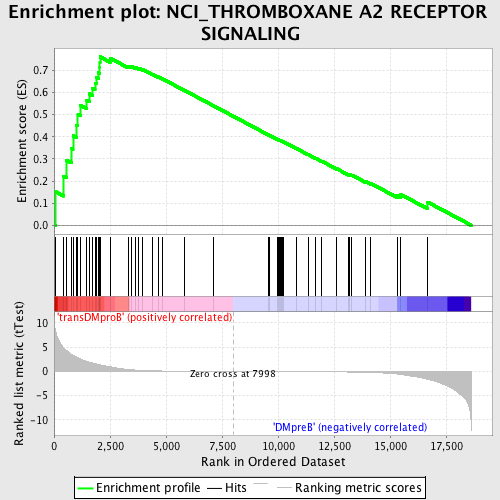
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
| GeneSet | NCI_THROMBOXANE A2 RECEPTOR SIGNALING |
| Enrichment Score (ES) | 0.7612875 |
| Normalized Enrichment Score (NES) | 1.7607701 |
| Nominal p-value | 0.0020533882 |
| FDR q-value | 0.037996117 |
| FWER p-Value | 0.029 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GNA15 | 19671 | 61 | 8.343 | 0.1518 | Yes | ||
| 2 | MAPK14 | 23313 | 429 | 4.797 | 0.2212 | Yes | ||
| 3 | PRKCH | 21246 | 560 | 4.274 | 0.2937 | Yes | ||
| 4 | LCK | 15746 | 786 | 3.491 | 0.3465 | Yes | ||
| 5 | ICAM1 | 19545 | 844 | 3.333 | 0.4054 | Yes | ||
| 6 | SLC9A3R1 | 11250 | 989 | 2.967 | 0.4528 | Yes | ||
| 7 | PRKCD | 21897 | 1065 | 2.795 | 0.5007 | Yes | ||
| 8 | GNG2 | 4790 | 1174 | 2.571 | 0.5427 | Yes | ||
| 9 | ADRBK1 | 23960 | 1453 | 2.060 | 0.5660 | Yes | ||
| 10 | RAB11A | 7013 | 1584 | 1.878 | 0.5939 | Yes | ||
| 11 | ARRB2 | 20806 | 1715 | 1.707 | 0.6187 | Yes | ||
| 12 | GNA13 | 20617 | 1827 | 1.605 | 0.6425 | Yes | ||
| 13 | ROCK1 | 5386 | 1898 | 1.517 | 0.6669 | Yes | ||
| 14 | GNB1 | 15967 | 1980 | 1.431 | 0.6892 | Yes | ||
| 15 | EGF | 15169 | 2044 | 1.359 | 0.7111 | Yes | ||
| 16 | VCAM1 | 5851 | 2047 | 1.358 | 0.7362 | Yes | ||
| 17 | PTGIR | 18376 | 2050 | 1.355 | 0.7613 | Yes | ||
| 18 | RAC1 | 16302 | 2500 | 0.969 | 0.7551 | No | ||
| 19 | ARHGEF1 | 1952 18343 3996 | 3312 | 0.395 | 0.7188 | No | ||
| 20 | TGM2 | 2657 | 3456 | 0.337 | 0.7174 | No | ||
| 21 | GNA12 | 9023 | 3615 | 0.280 | 0.7141 | No | ||
| 22 | PLCB2 | 5262 | 3787 | 0.233 | 0.7092 | No | ||
| 23 | FGR | 4723 | 3942 | 0.194 | 0.7045 | No | ||
| 24 | HCK | 14787 | 4391 | 0.120 | 0.6826 | No | ||
| 25 | GNA14 | 23907 | 4649 | 0.091 | 0.6705 | No | ||
| 26 | ARR3 | 24282 9305 | 4852 | 0.076 | 0.6610 | No | ||
| 27 | PRKG1 | 5289 | 5807 | 0.038 | 0.6103 | No | ||
| 28 | LYN | 16281 | 7117 | 0.012 | 0.5400 | No | ||
| 29 | FYN | 3375 3395 20052 | 9588 | -0.022 | 0.4074 | No | ||
| 30 | GNAQ | 4786 23909 3685 | 9625 | -0.023 | 0.4059 | No | ||
| 31 | PTGDR | 21866 | 9975 | -0.028 | 0.3876 | No | ||
| 32 | PRKCE | 9575 | 10004 | -0.029 | 0.3866 | No | ||
| 33 | NOS3 | 16906 885 | 10060 | -0.030 | 0.3842 | No | ||
| 34 | PRKCQ | 2873 2831 | 10099 | -0.030 | 0.3827 | No | ||
| 35 | YES1 | 5930 | 10146 | -0.031 | 0.3808 | No | ||
| 36 | GNB5 | 2985 9029 3153 | 10186 | -0.032 | 0.3793 | No | ||
| 37 | SELE | 14074 | 10217 | -0.032 | 0.3783 | No | ||
| 38 | DNM1 | 2777 8857 | 10837 | -0.043 | 0.3458 | No | ||
| 39 | SRC | 5507 | 11343 | -0.054 | 0.3196 | No | ||
| 40 | PRKCA | 20174 | 11681 | -0.062 | 0.3026 | No | ||
| 41 | PRKACA | 18549 3844 | 11953 | -0.071 | 0.2893 | No | ||
| 42 | EGFR | 1329 20944 | 12619 | -0.098 | 0.2553 | No | ||
| 43 | ADRBK2 | 11499 | 13153 | -0.129 | 0.2290 | No | ||
| 44 | AKT1 | 8568 | 13190 | -0.131 | 0.2295 | No | ||
| 45 | RHOA | 8624 4409 4410 | 13287 | -0.138 | 0.2269 | No | ||
| 46 | GNA11 | 3300 3345 19670 | 13911 | -0.196 | 0.1970 | No | ||
| 47 | MAPK11 | 2264 9618 22163 | 14126 | -0.222 | 0.1896 | No | ||
| 48 | PRKCZ | 5260 | 15328 | -0.546 | 0.1350 | No | ||
| 49 | SYK | 21636 | 15479 | -0.643 | 0.1389 | No | ||
| 50 | BLK | 3205 21791 | 16677 | -1.613 | 0.1044 | No |

